Advanced Mixed-Integer Programming (MIP) Example Using CVXPY
Problem: Facility Layout Problem with Sequencing and Capacity Constraints
The Facility Layout Problem is a complex optimization problem where multiple facilities need to be arranged in a layout to minimize operational costs, such as transportation between facilities, while respecting various constraints like sequencing, capacity, and adjacency requirements.
This problem becomes more challenging when sequencing and capacity constraints are introduced, turning it into a non-trivial MIP problem.
Problem Setup
Objective:
The goal is to place facilities on a grid layout to minimize the total transportation cost, considering:
- Sequencing requirements (some facilities must be placed in a specific order).
- Capacity constraints (some facilities can only be placed in specific regions due to space limitations).
Variables:
- $(x_{ij})$: Binary variable that equals $1$ if facility $(i)$ is placed in position $(j)$, $0$ otherwise.
- $(y_{ij})$: Binary variable that equals $1$ if facility $(i)$ precedes facility $(j)$ in the sequence.
Parameters:
- $(c_{ij})$: Cost of placing facility $(i)$ next to facility $(j)$.
- $(p_i)$: Capacity requirement for each facility.
- $(g_j)$: Capacity available at each grid position.
Constraints:
- Unique Placement: Each facility must be placed in exactly one position.
- No Overlap: Each position can hold at most one facility.
- Sequencing: Facilities must follow a predefined sequence if required.
- Capacity: Facilities can only be placed in positions that meet their capacity requirements.
Mathematical Formulation
Objective:
$$
\text{Minimize} \quad \sum_{i} \sum_{j} c_{ij} x_{ij}
$$Constraints:
- Each facility is placed in exactly one position:
$$
\sum_{j} x_{ij} = 1, \quad \forall i
$$ - Each position can hold at most one facility:
$$
\sum_{i} x_{ij} \leq 1, \quad \forall j
$$ - Sequencing constraints between facilities:
$$
y_{ij} + y_{ji} = 1, \quad \forall i \neq j
$$ - Capacity constraints:
$$
p_i \times x_{ij} \leq g_j, \quad \forall i, j
$$
- Each facility is placed in exactly one position:
Solving with CVXPY
Here’s a detailed implementation using $CVXPY$:
1 | import cvxpy as cp |
Explanation of the Code:
Decision Variables:
x[i, j]: Binary variable indicating whether facility $( i )$ is placed in position $( j )$.y[i, j]: Binary variable indicating whether facility $( i )$ precedes facility $( j )$ in the sequence.
Objective:
- The objective is to minimize the total placement cost, computed by multiplying the placement matrix with the cost matrix.
Constraints:
- Each facility must be assigned exactly one position, and each position can hold at most one facility.
- Sequencing constraints ensure that the required order between facilities is maintained.
- Capacity constraints enforce that a facility can only be placed in a position with sufficient available capacity.
Solver:
- The problem is solved using
GLPK_MI, an open-source solver suitable for MIP problems.
- The problem is solved using
Output:
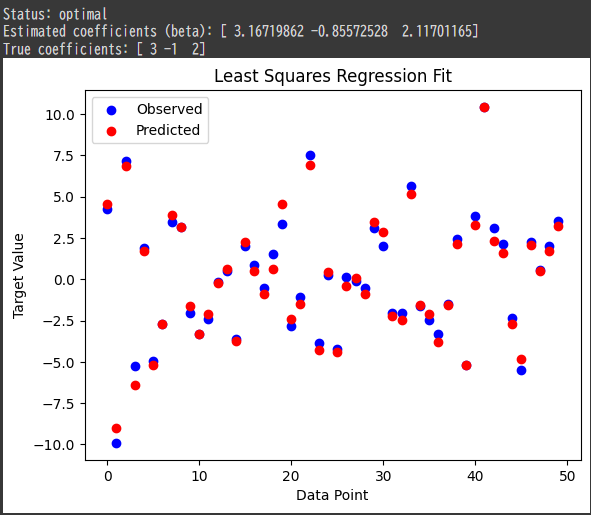
1 | Status: optimal |
Conclusion:
This advanced Mixed-Integer Programming (MIP) example demonstrates how to solve a complex facility layout problem with sequencing and capacity constraints using $CVXPY$.
The use of binary variables allows us to model the discrete nature of facility placements and sequencing requirements, making MIP a powerful tool for such complex layout optimization problems.