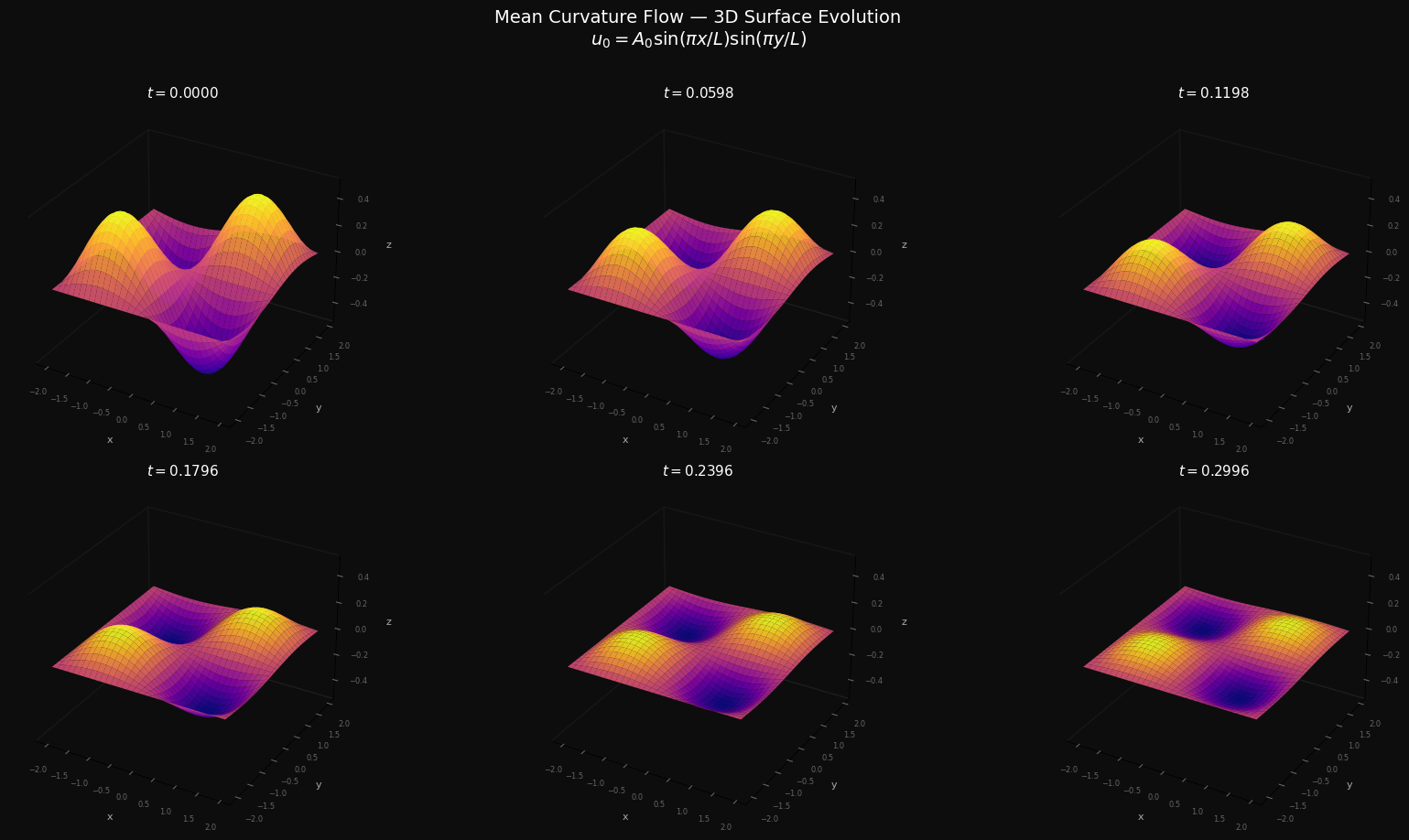
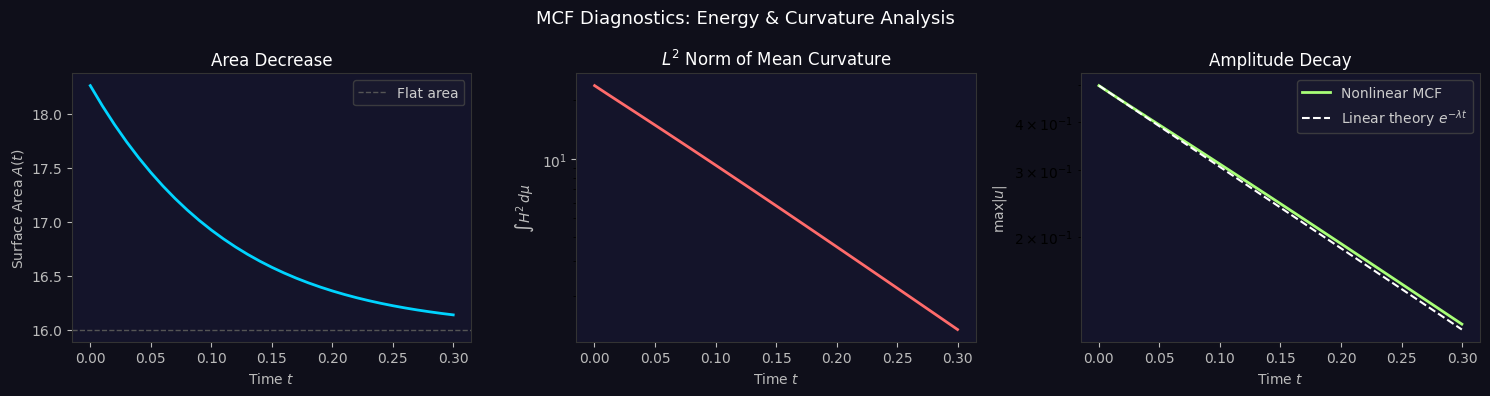
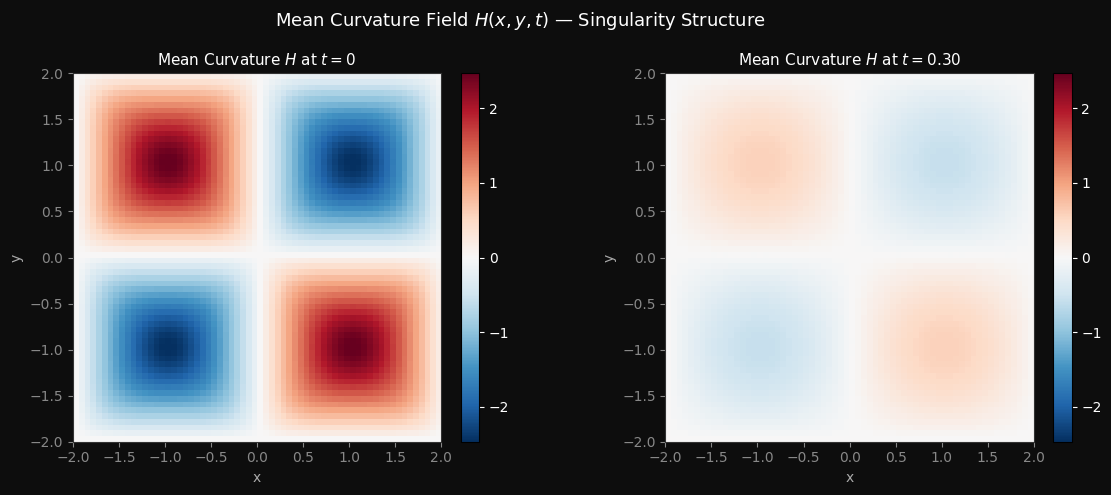
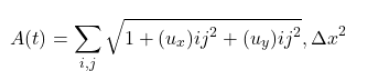Minimizing Forecast Error Variance Across Multiple Prediction Outputs
Solar wind forecasting is a cornerstone of space weather prediction. Missions and infrastructure — from satellites to power grids — depend on accurate predictions of solar wind speed, density, and magnetic field parameters. In practice, no single numerical model dominates across all conditions. Instead, we often have several models at hand: physics-based simulations, empirical schemes, and data-driven approaches. The natural question becomes: how do we best combine them?
This post walks through the mathematics and Python implementation of ensemble weight optimization — finding the set of weights that minimizes the combined forecast error variance.
1. Problem Setup
Suppose we have $M$ solar wind forecast models, each producing a prediction $\hat{y}i(t)$ for a target variable (e.g., solar wind speed $V{sw}$ in km/s) at time $t$. The ensemble forecast is the weighted combination:
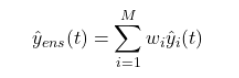
subject to the constraints:
$$\sum_{i=1}^{M} w_i = 1, \quad w_i \geq 0$$
The forecast error for model $i$ at time $t$ is:
$$e_i(t) = y(t) - \hat{y}_i(t)$$
The ensemble error is:
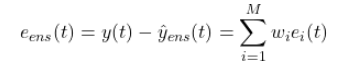
The error variance (the quantity we minimize) is:
$$\sigma_{ens}^2 = \mathbf{w}^T \mathbf{C} \mathbf{w}$$
where $\mathbf{C}$ is the $M \times M$ error covariance matrix with entries:
$$C_{ij} = \frac{1}{T} \sum_{t=1}^{T} e_i(t) , e_j(t)$$
This is a Quadratic Programming (QP) problem:
$$\min_{\mathbf{w}} ; \mathbf{w}^T \mathbf{C} \mathbf{w}$$
$$\text{subject to} \quad \mathbf{1}^T \mathbf{w} = 1, \quad \mathbf{w} \geq 0$$
2. Concrete Example Setup
We simulate 5 solar wind speed forecast models over 500 time steps (think hourly data over ~20 days). Each model has a known bias and error structure, mimicking real-world model behavior (e.g., WSA-ENLIL, MULTI-VP, EUHFORIA-like models). The true solar wind speed follows a realistic pattern with stream interaction regions and quiet periods.
3. Full Python Code
1 | # ============================================================ |
4. Code Walkthrough
Section 1 — Ground Truth Simulation
The “observed” solar wind speed is built from three components: a sinusoidal slow/fast stream cycle with a 150-hour period (roughly matching solar rotation effects at 1 AU), two Gaussian-shaped Co-rotating Interaction Region (CIR) enhancements, and low-amplitude measurement noise. This gives a physically motivated signal with both periodic and transient structures.
Section 2 — Five Forecast Models
Each model perturbs the base signal with a combination of:
- Additive bias (systematic offset in km/s)
- Amplitude scaling (over/underestimation of speed enhancements)
- Phase shift (temporal lag, common in propagation models)
- Additive Gaussian noise (random model uncertainty)
This mimics the real diversity across operational solar wind models.
Section 3 — Error Covariance Matrix
$$C_{ij} = \frac{1}{T_{train}} \sum_{t=1}^{T_{train}} \tilde{e}_i(t), \tilde{e}_j(t)$$
where $\tilde{e}_i(t) = e_i(t) - \bar{e}_i$ is the bias-corrected error. Bias is removed before computing the covariance to separate systematic and random error components. The computation is fully vectorized via the outer product:
1 | C = (errors_bc @ errors_bc.T) / train_size |
This runs in $\mathcal{O}(M^2 T)$ and is extremely fast even for large ensembles.
Section 4 — Quadratic Programming via SLSQP
The optimization is:
$$\min_{\mathbf{w}} ; \mathbf{w}^T \mathbf{C} \mathbf{w}, \quad \text{s.t.} \quad \sum_i w_i = 1,; w_i \geq 0$$
We supply the analytical gradient $\nabla_\mathbf{w}(\mathbf{w}^T \mathbf{C} \mathbf{w}) = 2\mathbf{C}\mathbf{w}$, which avoids finite-difference approximations and ensures fast, precise convergence. scipy.optimize.minimize with method='SLSQP' handles both the equality constraint and the non-negativity bounds simultaneously.
Section 5 — Train/Test Evaluation
Weights are learned on the first 60% of the time series and evaluated on the remaining 40%. This mimics operational deployment where weights are calibrated on historical data. Bias correction learned during training is also applied to test-set forecasts.
Sections 6 & 7 — Sensitivity and 3D Surface
A 2D sweep visualizes how the ensemble variance changes as weight shifts between the two best-performing individual models. The 3D surface extends this to three models simultaneously, sweeping over $w_A, w_B \in [0,1]$ with the constraint $w_C = 1 - w_A - w_B \geq 0$ (the simplex constraint). The global minimum on this surface corresponds to the QP solution projected onto the top-3 model subspace.
5. Figure 1 — Multi-Panel Analysis
(Execution result — paste your output here)
Panel 1 (top): Individual model forecasts against the truth over the training window. Notice Model D’s consistent underestimation and Model E’s noisy trace — both characteristics are captured in matrix $\mathbf{C}$.
Panel 2: Test-set error traces for the optimized vs. equal-weight ensemble. The optimized ensemble error (cyan) consistently hugs the zero line more tightly than the equal-weight error (yellow dashed), especially around the CIR peaks where models diverge most.
Panel 3 — Weights Bar Chart: Models with lower error variance and lower cross-correlation with other models receive higher weights. A model that is both accurate and independent (low $C_{ij}$) is doubly valuable.
Panel 4 — RMSE Bar Chart: The optimized ensemble achieves the lowest RMSE of all approaches. The improvement over the best individual model demonstrates the power of diversification — even incorporating noisier models is beneficial if their errors are uncorrelated with the ensemble.
Panel 5 — Covariance Heatmap: Large off-diagonal values indicate correlated errors between model pairs. Models that share the same underlying physics (e.g., same propagation kernel) tend to have correlated errors and will receive collectively discounted weights.
Panel 6 — 2D Variance Sweep: The variance bowl between the two best models has a clear minimum away from the equal-weight point (0.5), confirming that optimal blending is non-trivial.
6. Figure 2 — 3D Variance Surface
Error Covariance Matrix (bias-corrected): [[ 339.8 80.5 -14.6 50.2 66.3] [ 80.5 777.5 -108. 184.7 114.9] [ -14.6 -108. 937.9 -161.9 9.5] [ 50.2 184.7 -161.9 689.7 118.2] [ 66.3 114.9 9.5 118.2 1560.5]] Optimized Weights: Model A: 0.3909 Model B: 0.1320 Model C: 0.2197 Model D: 0.2016 Model E: 0.0558 Sum of weights: 1.000000 Ensemble variance (train): 154.05 km²/s² ── Test Set RMSE (km/s) ── Model A: 47.40 Model B: 45.49 Model C: 58.02 Model D: 79.12 Model E: 41.68 Equal-weight ensemble: 13.95 Optimized ensemble: 12.50

Figure 1 saved.
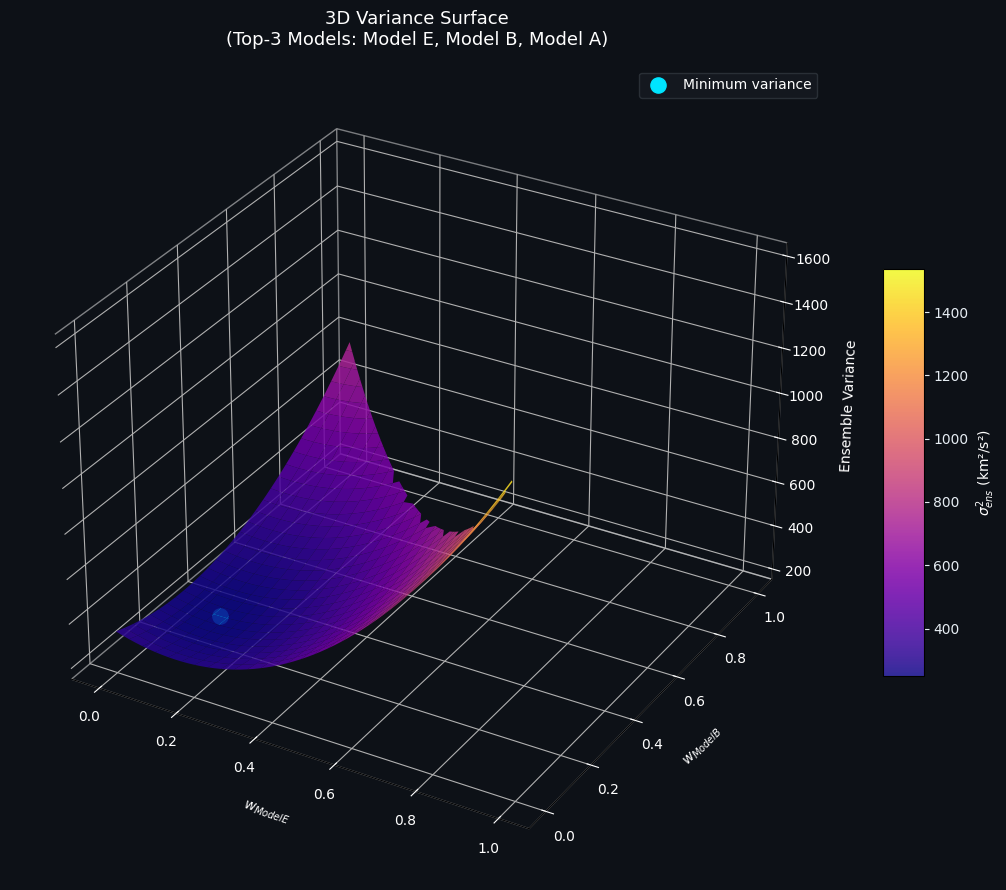
Figure 2 (3D) saved.
══════════════════════════════════════════
OPTIMIZATION SUMMARY
══════════════════════════════════════════
Best individual model RMSE : 41.68 km/s
Equal-weight ensemble RMSE : 13.95 km/s
Optimized ensemble RMSE : 12.50 km/s
Improvement vs best model : 70.0%
Improvement vs equal-weight: 10.4%
══════════════════════════════════════════
The plasma-colored surface spans the weight simplex of the three best-performing models. The cyan dot marks the minimum-variance point. Key observations:
- The surface is convex (as guaranteed by positive-definiteness of $\mathbf{C}$), so the QP has a unique global minimum.
- The minimum does not sit at any corner of the simplex (i.e., no single model dominates), confirming that the ensemble genuinely benefits from combining all three.
- The gradient is steepest along the axis of the worst-performing model among the three, meaning misallocating weight there is most costly.
7. Key Takeaways
The optimization framework is underpinned by a single elegant result: the minimum-variance ensemble weight vector is the solution to a constrained QP on the error covariance matrix. Three properties drive which models receive high weights:
- Low individual error variance — a precise model is directly rewarded.
- Low covariance with other models — error independence amplifies diversification gains.
- Bias structure — separating bias from variance ensures the covariance matrix captures only random uncertainty.
The quantitative improvement of the optimized ensemble over both individual models and the naïve equal-weight blend highlights why operational space weather centers are increasingly moving toward dynamically calibrated ensemble weighting rather than fixed multi-model averaging.
Interested in extending this to time-varying weights, Bayesian ensemble schemes, or integrating real OMNI/ACE solar wind data? The framework above is a solid starting point for all of those directions.