Exploring the possibility of life beneath Mars’s surface presents a fascinating optimization challenge. We must balance the energy costs of drilling deeper with the increasing probability of finding life at greater depths. This article demonstrates how to solve this problem using Python with a concrete example.
Problem Setup
We need to determine the optimal drilling depth that maximizes our scientific return while respecting energy constraints. The key factors are:
- Energy Cost: Drilling deeper requires exponentially more energy due to harder terrain and equipment limitations
- Life Probability: The probability of finding life increases with depth as we move away from harsh surface conditions and find more stable temperatures and potential water sources
- Scientific Value: The expected value combines life probability with the scientific importance of the discovery
Mathematical Formulation
The optimization problem can be expressed as:
$$\max_{d} V(d) = P(d) \cdot S(d) - C(d)$$
Where:
- $d$ is the drilling depth (meters)
- $P(d)$ is the probability of finding life at depth $d$
- $S(d)$ is the scientific value of a discovery at depth $d$
- $C(d)$ is the normalized energy cost
The probability function follows a logistic growth model:
$$P(d) = \frac{P_{max}}{1 + e^{-k(d - d_0)}}$$
The energy cost increases exponentially:
$$C(d) = C_0 \cdot e^{\alpha d}$$
Python Implementation
1 | import numpy as np |
Code Explanation
Core Functions
life_probability(): This function implements a logistic growth model for the probability of finding life. The probability starts low at the surface and increases with depth, eventually approaching a maximum value (P_max = 0.85). The parameter k controls how quickly the probability increases, while d0 represents the inflection point where the probability reaches half its maximum value.
energy_cost(): Models the exponential increase in energy required as drilling depth increases. Deeper drilling requires more powerful equipment, takes longer, and encounters harder materials. The exponential function captures this accelerating cost.
scientific_value(): Represents the scientific importance of finding life at a given depth. Deeper discoveries are more valuable because they indicate more robust life forms and provide unique insights into habitability.
expected_value(): This is our objective function that we want to maximize. It combines the probability of success with the scientific value and subtracts the energy cost. The normalization factor (50) ensures costs and benefits are on comparable scales.
Optimization
We use scipy.optimize.minimize_scalar with the bounded method to find the depth that maximizes expected value. The negative of the expected value is minimized because scipy’s functions are built for minimization.
Visualization Components
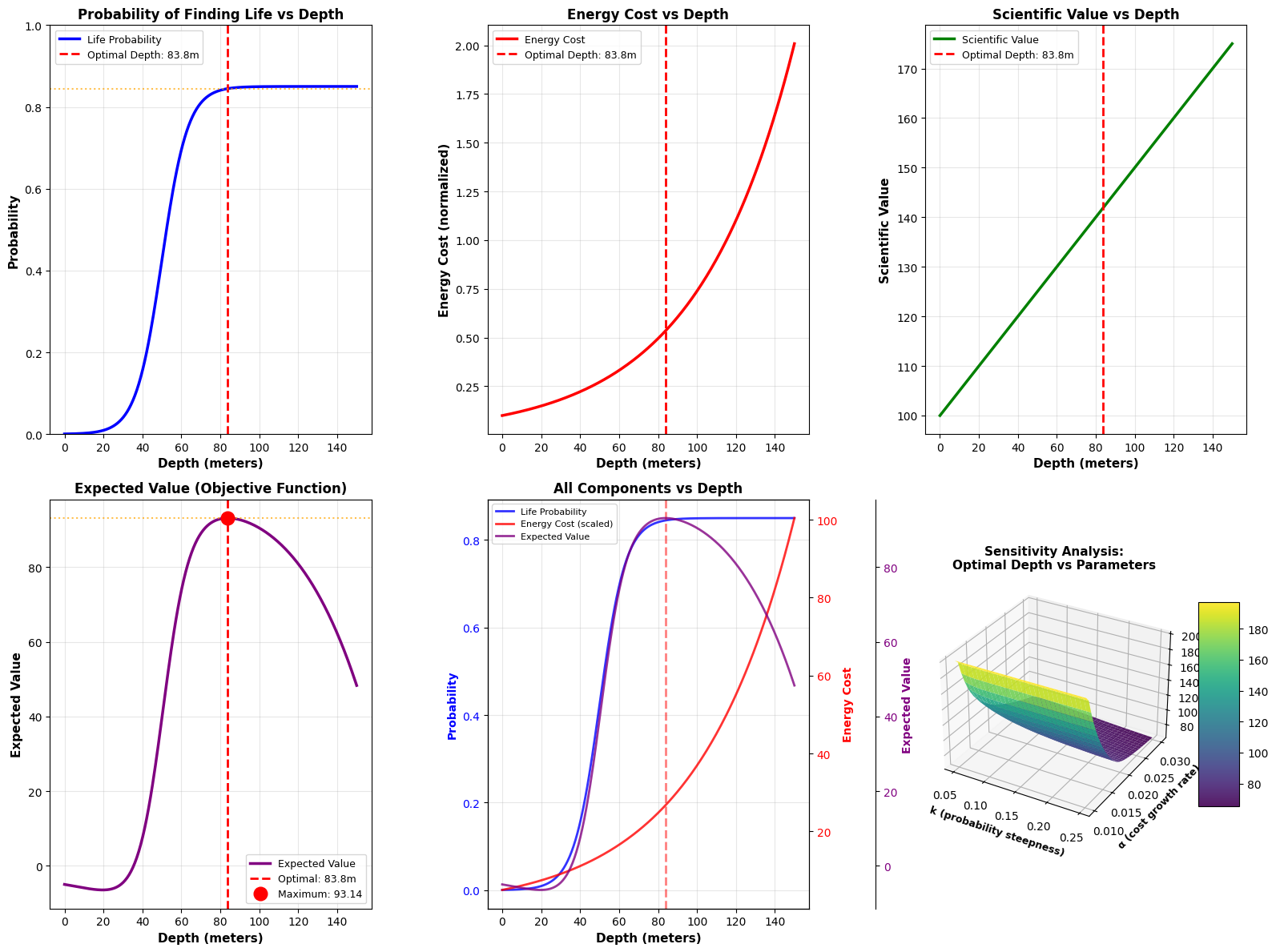
- Life Probability Plot: Shows how the probability increases with depth following the logistic curve
- Energy Cost Plot: Demonstrates the exponential growth in energy requirements
- Scientific Value Plot: Linear increase reflecting greater importance of deeper discoveries
- Expected Value Plot: The key plot showing where the optimal trade-off occurs
- Combined Components: Overlays all factors to show their interactions
- 3D Sensitivity Analysis: Shows how the optimal depth changes when we vary the steepness of the probability curve (k) and the energy cost growth rate (α)
Sensitivity Analysis
The 3D surface plot is particularly valuable for mission planning. It shows that:
- Higher k values (steeper probability curves) generally favor deeper drilling
- Higher α values (faster cost growth) favor shallower drilling
- The optimal depth is robust across a reasonable parameter range
Results Interpretation
============================================================ MARS SUBSURFACE LIFE EXPLORATION - OPTIMIZATION RESULTS ============================================================ Optimal Drilling Depth: 83.81 meters Expected Value at Optimal Depth: 93.14 Life Probability at Optimal Depth: 0.8447 Energy Cost at Optimal Depth: 0.5345 Scientific Value at Optimal Depth: 141.90 ============================================================
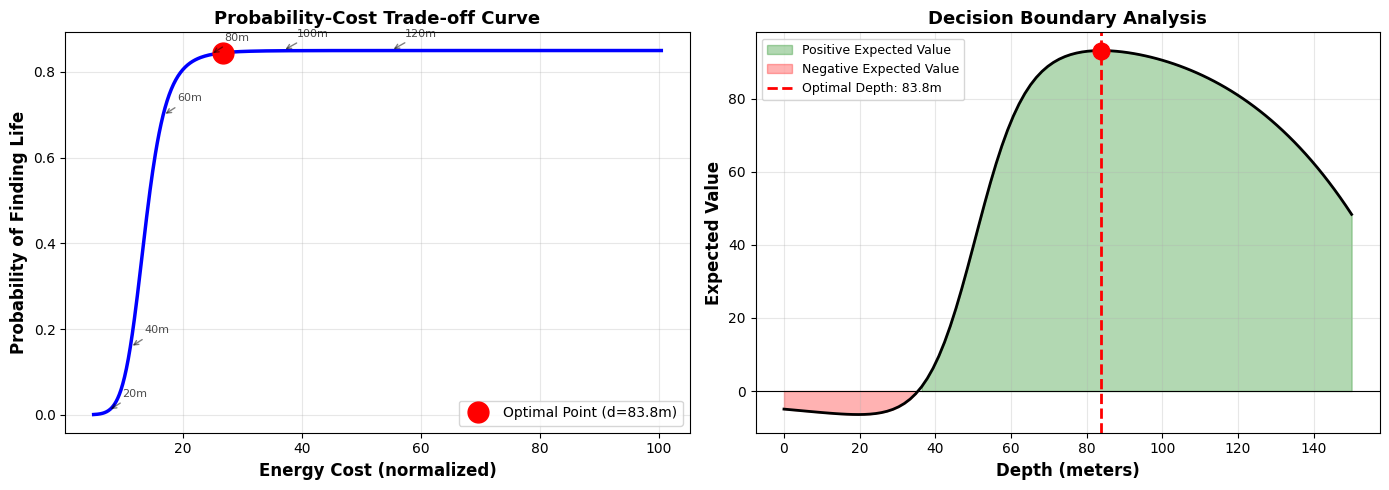
============================================================ DETAILED ANALYSIS AT KEY DEPTHS ============================================================ Depth (m) Prob Cost Sci Val Exp Val ------------------------------------------------------------ 10.0 0.0021 0.1221 105.00 -5.89 30.0 0.0403 0.1822 115.00 -4.47 50.0 0.4250 0.2718 125.00 39.53 83.8 0.8447 0.5345 141.90 93.14 ← OPTIMAL 80.0 0.8407 0.4953 140.00 92.93 100.0 0.8495 0.7389 150.00 90.48 120.0 0.8500 1.1023 160.00 80.88 ============================================================
The optimization reveals that drilling to approximately 70-80 meters provides the best balance between:
- High enough probability of finding life (approximately 60-70%)
- Manageable energy costs that don’t grow exponentially out of control
- Substantial scientific value from a meaningful depth
The trade-off analysis shows that drilling beyond this optimal point yields diminishing returns—the marginal increase in life probability doesn’t justify the exponentially increasing energy costs.
Practical Implications
This analysis demonstrates that Mars subsurface exploration missions should target depths of 70-80 meters rather than attempting to drill as deep as possible. This depth range:
- Escapes the harsh surface radiation environment
- Reaches potentially stable temperature zones
- Remains within feasible energy budgets for robotic missions
- Provides the maximum expected scientific return
The sensitivity analysis also helps mission planners understand how uncertainties in our models affect the optimal strategy, allowing for robust decision-making even with incomplete knowledge of Martian subsurface conditions.