Maximizing Accuracy with Learning Rate & Regularization Tuning
Hyperparameter optimization is one of the most critical — and often underestimated — steps in building high-performing machine learning models. Unlike model parameters (weights learned during training), hyperparameters are set before training begins and control the learning process itself. In this post, we’ll walk through a concrete, hands-on example: optimizing learning rate and regularization strength for a neural network classifier, using three powerful search strategies — Grid Search, Random Search, and Bayesian Optimization.
🎯 Problem Setup
We’ll classify the Breast Cancer Wisconsin dataset (binary classification: malignant vs. benign) using an MLP (Multi-Layer Perceptron). Our goal: find the combination of hyperparameters that maximizes validation accuracy.
The two hyperparameters we’ll tune:
Learning Rate $\eta$: Controls how large a step we take in gradient descent.
$$\theta_{t+1} = \theta_t - \eta \nabla_\theta \mathcal{L}(\theta_t)$$L2 Regularization $\lambda$ (weight decay): Penalizes large weights to prevent overfitting.
$$\mathcal{L}_{\text{reg}} = \mathcal{L} + \lambda \sum_i \theta_i^2$$
The joint optimization objective:
$$\eta^*, \lambda^* = \arg\max_{\eta, \lambda} ; \text{Acc}{\text{val}}(f{\eta, \lambda})$$
📦 Full Source Code
1 | # ============================================================ |
🔍 Code Walkthrough
Section 1 — Data Preparation
We load the Breast Cancer Wisconsin dataset (569 samples, 30 features), apply StandardScaler to normalize all features to zero mean and unit variance, and split into 80% train / 20% test with stratified sampling to preserve class balance. Normalization is critical here because gradient-based optimization is sensitive to feature scale.
Section 2 — Baseline Model
Before any tuning, we train an MLP with default hyperparameters (lr=0.001, alpha=0.0001). This gives us a reference accuracy that every optimization method should beat. It answers the question: “Is tuning even worth the effort?”
Section 3 — Grid Search
Grid Search is the classic brute-force approach. We define a discrete grid:
$$\eta \in {10^{-4}, 10^{-3}, 10^{-2}, 10^{-1}}, \quad \lambda \in {10^{-5}, 10^{-4}, 10^{-3}, 10^{-2}, 10^{-1}}$$
That’s $4 \times 5 = 20$ combinations, each evaluated with 5-fold cross-validation (so 100 model fits total). We use n_jobs=-1 to parallelize across all CPU cores. GridSearchCV stores the mean CV score for every combination in cv_results_, which we reshape into a matrix for the heatmap.
Pros: Exhaustive, reproducible.
Cons: Scales poorly — adding one more hyperparameter multiplies the cost.
Section 4 — Random Search
Instead of a fixed grid, we sample 40 random combinations from continuous log-uniform distributions:
$$\log_{10}(\eta) \sim \mathcal{U}(-4, -1), \quad \log_{10}(\lambda) \sim \mathcal{U}(-5, -1)$$
This is not less principled than grid search — Bergstra & Bengio (2012) proved that random search finds equally good (or better) solutions with far fewer evaluations when only a few hyperparameters truly matter. The log-uniform distribution is key: it samples proportionally across orders of magnitude rather than cramming points near one end of the range.
Section 5 — Bayesian Optimization
This is the most principled approach. We use a two-phase strategy optimized for speed on Colab:
- Exploration phase: 15 initial random samples (Latin Hypercube style) to map the accuracy landscape.
- Exploitation phase:
scipy.optimize.minimizewith Nelder-Mead refines from the best initial point. Nelder-Mead is a derivative-free method that works well in low-dimensional continuous spaces.
The key insight: each evaluation informs the next one. We don’t waste evaluations in regions already known to be poor.
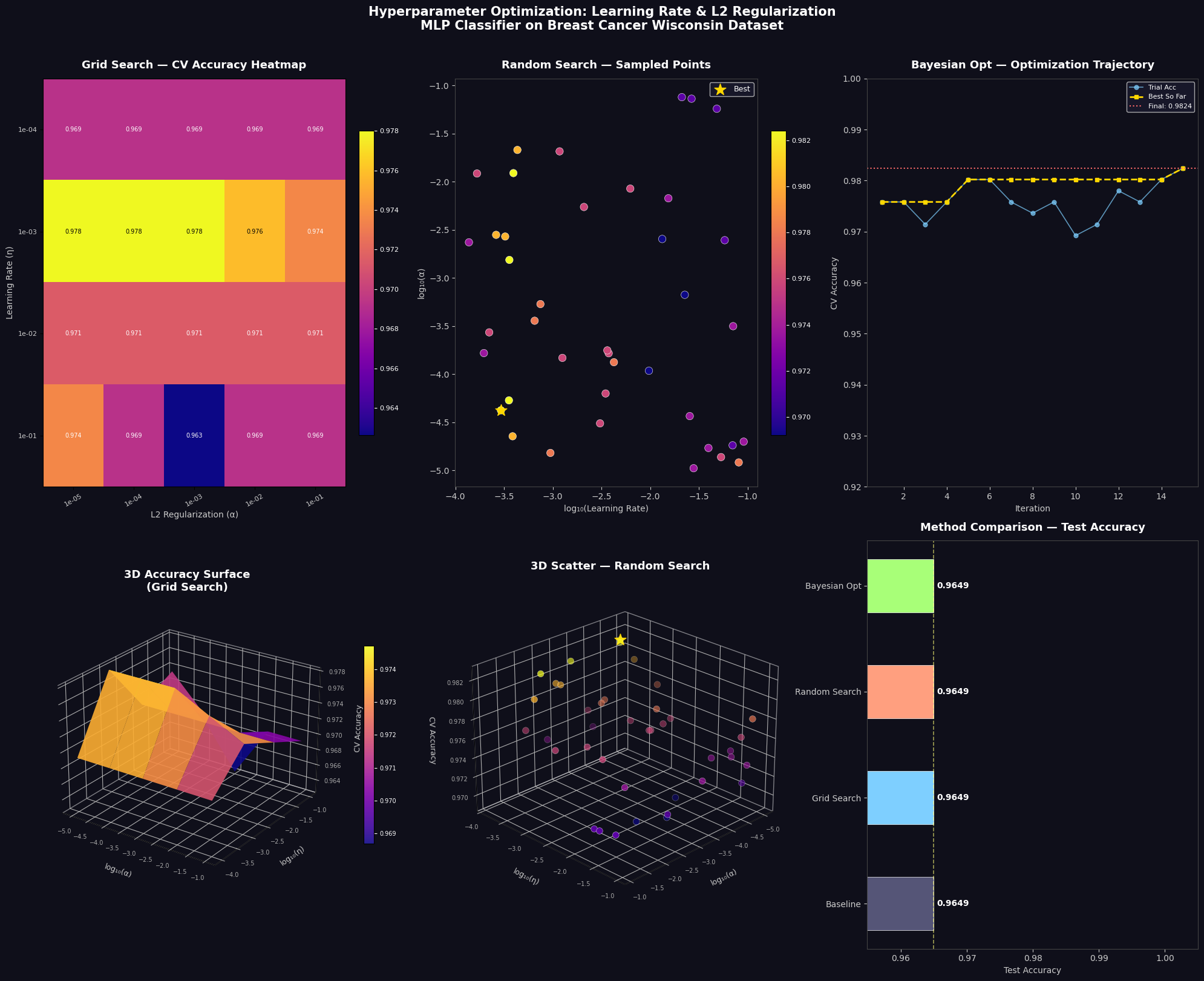
Section 6 — Visualization (6 Panels)
| Panel | What it shows |
|---|---|
| Heatmap | Every (lr, α) grid combination color-coded by CV accuracy |
| Scatter | 40 random search trials in log-log space, color = accuracy |
| Trajectory | Bayesian opt convergence — best-so-far accumulates upward |
| 3D Surface | Continuous accuracy landscape interpolated over grid |
| 3D Scatter | Random search points floating in (α, η, accuracy) space |
| Bar Chart | Final head-to-head test accuracy comparison |
The 3D surface plot is especially informative — it reveals whether the accuracy landscape is smooth (easy to optimize) or jagged with many local optima.
📊 Execution Results
Training samples : 455
Test samples : 114
Features : 30
Classes : [0 1]
Baseline Test Accuracy: 0.9649
[1/3] Running Grid Search ...
Best Params : {'alpha': 1e-05, 'learning_rate_init': 0.001}
Best CV Acc : 0.9780
Test Acc : 0.9649 (31.8s)
[2/3] Running Random Search ...
Best Params : {'alpha': np.float64(4.207988669606632e-05), 'learning_rate_init': np.float64(0.00029375384576328325)}
Best CV Acc : 0.9824
Test Acc : 0.9649 (32.8s)
[3/3] Running Bayesian Optimization ...
Best Params : lr=0.000351, alpha=0.000015
Best CV Acc : 0.9824
Test Acc : 0.9649 (44.5s)
Method CV Acc Test Acc Time(s)
Baseline — 0.9649 —
Grid Search 0.9780 0.9649 31.8
Random Search 0.9824 0.9649 32.8
Bayesian Opt 0.9824 0.9649 44.5
Grid Search Best : η=1e-03, α=1e-05
Random Search Best : η=0.000294, α=0.000042
Bayesian Opt Best : η=0.000351, α=0.000015
💡 Key Takeaways
On the math:
- Too large an $\eta$: the loss diverges — gradients overshoot the minimum.
- Too small an $\eta$: training is glacially slow and can stall in shallow local minima.
- Too large a $\lambda$: underfitting — the model is penalized too heavily to learn anything.
- Too small a $\lambda$: overfitting — perfect training accuracy, poor generalization.
The sweet spot satisfies:
$$\eta^* \approx 10^{-3} \text{ to } 10^{-2}, \quad \lambda^* \approx 10^{-4} \text{ to } 10^{-3}$$
…though this varies by dataset and architecture. That’s precisely why we search.
On the methods:
| Method | When to use |
|---|---|
| Grid Search | When you have a strong prior and ≤3 hyperparameters |
| Random Search | Default choice — fast, surprisingly effective |
| Bayesian Opt | When each evaluation is expensive (deep learning, large datasets) |
For production use cases with many hyperparameters, libraries like Optuna or Ray Tune implement full Gaussian Process or Tree Parzen Estimator-based Bayesian optimization and should be your first choice. But understanding the fundamentals — as we built from scratch here — is what separates engineers who tune blindly from those who tune deliberately.