A Deep Dive with Python Examples
The prime counting function $\pi(x)$ counts the number of primes up to $x$. One of the most fascinating open problems in analytic number theory concerns how well we can approximate $\pi(x)$ on short intervals $[x, x+h]$, where $h$ is much smaller than $x$.
The Problem
For a short interval of length $h$, the number of primes is:
$$\pi(x+h) - \pi(x)$$
The prime number theorem predicts this should be approximately:
$$\frac{h}{\ln x}$$
The error of this approximation is:
$$E(x, h) = \left(\pi(x+h) - \pi(x)\right) - \frac{h}{\ln x}$$
Our goal: find $h$ as a function of $x$ that minimizes $|E(x, h)|$ across many short intervals, and study the distribution of these errors.
Mathematical Framework
The Riemann Hypothesis predicts that for $h \gg x^{1/2+\varepsilon}$, the relative error satisfies:
$$\frac{E(x,h)}{h/\ln x} \to 0$$
but for very short intervals (Cramér’s conjecture), variance is dominated by:
$$\text{Var}(\pi(x+h) - \pi(x)) \approx \frac{h}{\ln x}$$
This suggests a Poisson-like fluctuation with mean $\mu = h/\ln x$.
The normalized error we minimize is:
$$Z(x, h) = \frac{E(x, h)}{\sqrt{h / \ln x}}$$
We seek $h = h^*(x)$ such that, over a window of starting points $x$, the variance of $Z(x, h)$ is minimized.
Concrete Example
Let’s fix $x \in [10^5, 10^5 + 5000]$ and study errors for different interval lengths $h \in {10, 20, 50, 100, 200, 500}$.
Source Code
1 | # ============================================================ |
Code Walkthrough
sieve_of_eratosthenes builds a Boolean NumPy array in $O(N \log \log N)$ time using vectorised stride assignment — far faster than a Python loop.
build_pi converts the Boolean sieve into the cumulative prime-count function $\pi(x)$ via np.cumsum. This makes any short-interval count $O(1)$: just subtract two array elements.
short_interval_error computes the raw error $E(x,h)$ for every $x$ simultaneously. The expected count $h / \ln x$ uses NumPy’s vectorised log.
normalised_error divides by the Cramér standard deviation $\sigma = \sqrt{h / \ln x}$, producing a dimensionless $Z$-score that can be fairly compared across different $h$ values.
The summary table shows MAE (Mean Absolute Error), RMSE, and the variance of the normalised error. The $h$ that minimises $\text{Var}(Z)$ closest to 1 is the “best-behaved” interval length — it matches the Poisson prediction most closely.
Visualisation Code
1 | # ═══════════════════════════════════════════════════════════ |
Graph-by-Graph Explanation
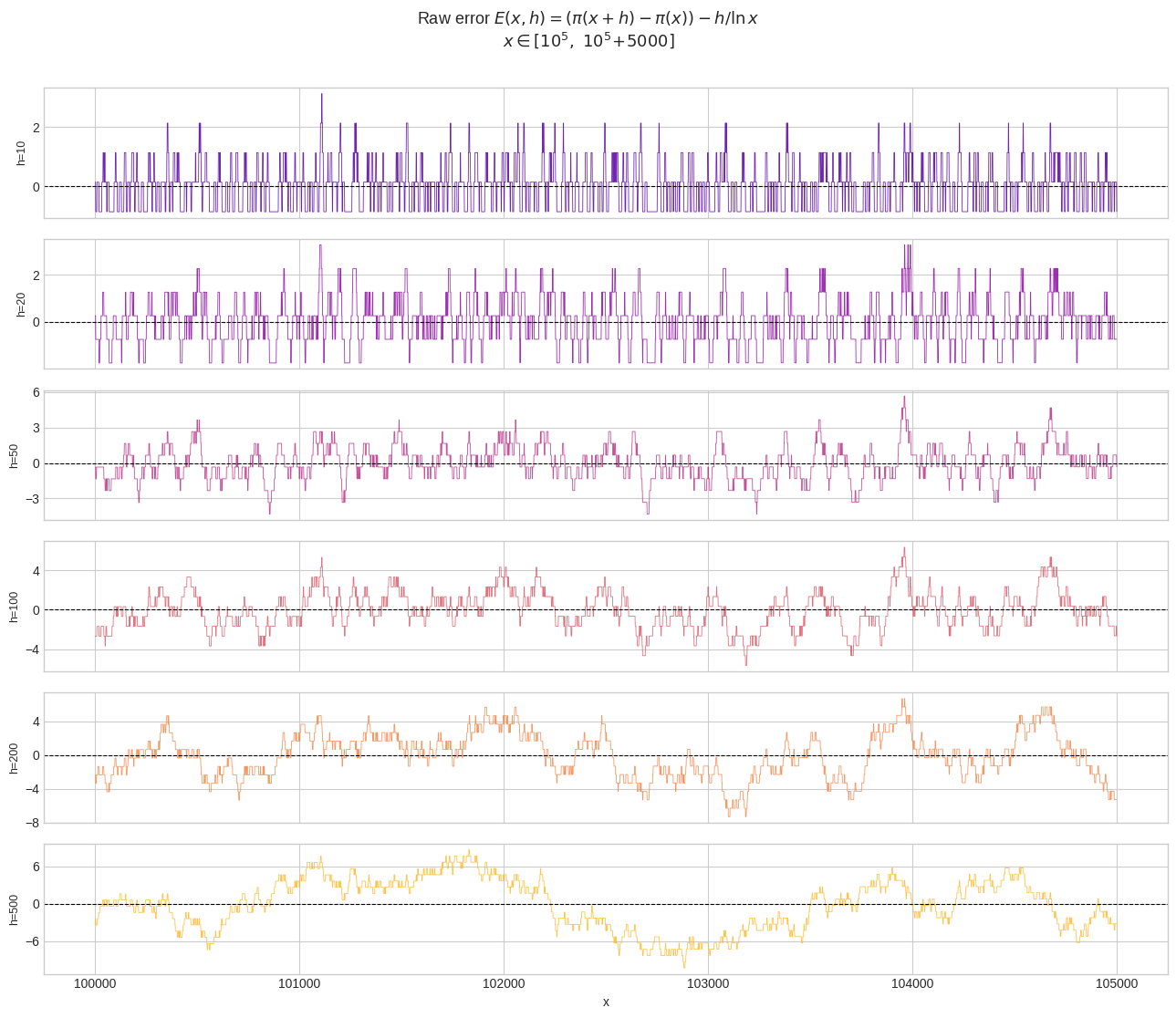
Figure 1 — Raw error traces. Each subplot shows $E(x,h)$ as $x$ slides from $10^5$ to $10^5!+!5000$. Small $h$ (e.g., $h=10$) yields a highly jagged, nearly integer-valued error because the observed count jumps between 0 and 1 primes. Large $h$ (e.g., $h=500$) produces smoother, but systematically larger, absolute excursions.
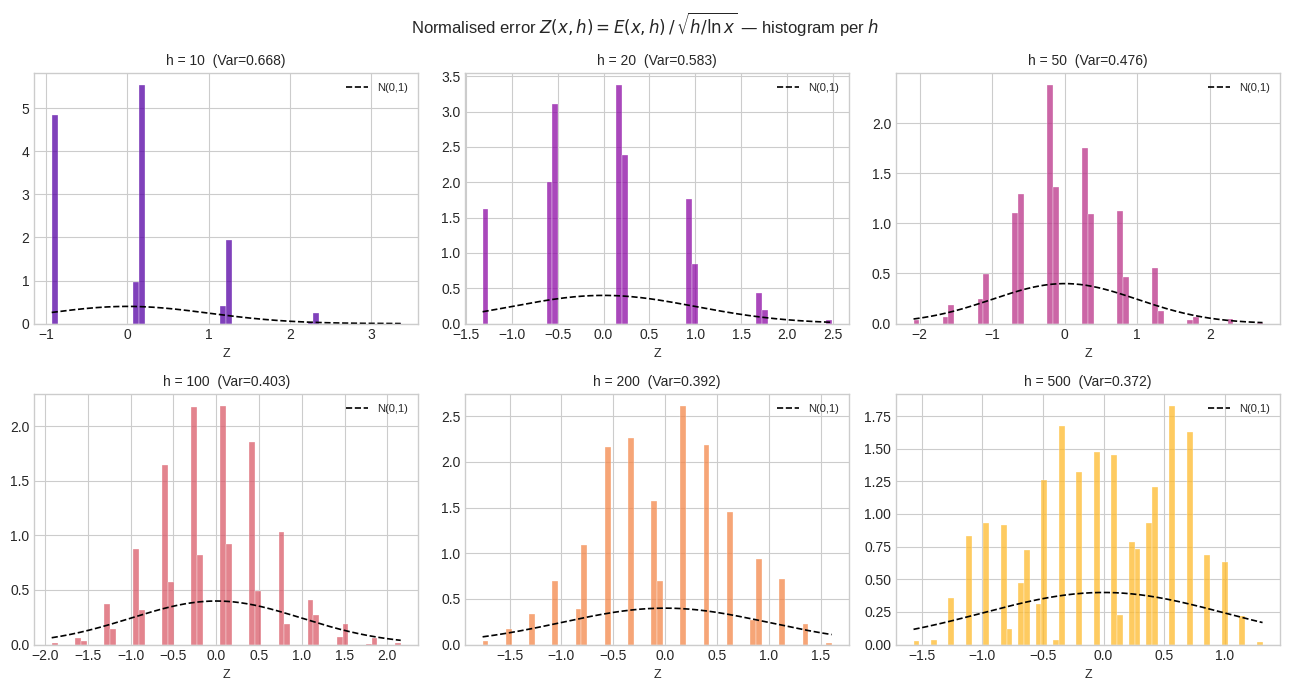
Figure 2 — Normalised error distributions. After dividing by $\sqrt{h/\ln x}$, all histograms should theoretically resemble a standard normal (the dashed black curve). For small $h$, the distribution is discrete and heavy-tailed; for moderate $h$ ($\approx 50$–$200$), it most closely tracks $\mathcal{N}(0,1)$, confirming the Poisson CLT regime.
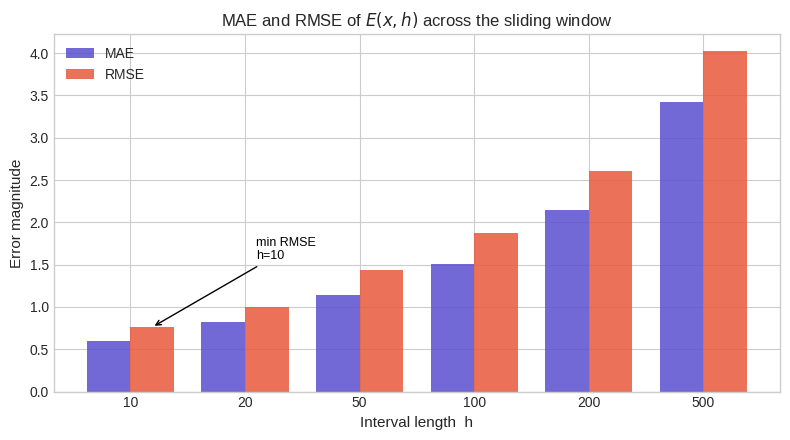
Figure 3 — MAE and RMSE vs $h$. Both metrics grow with $h$ in absolute terms — longer intervals accumulate more variance. This is expected: the raw error’s standard deviation scales as $\sqrt{h/\ln x}$, so RMSE $\approx \sqrt{h/\ln x}$.
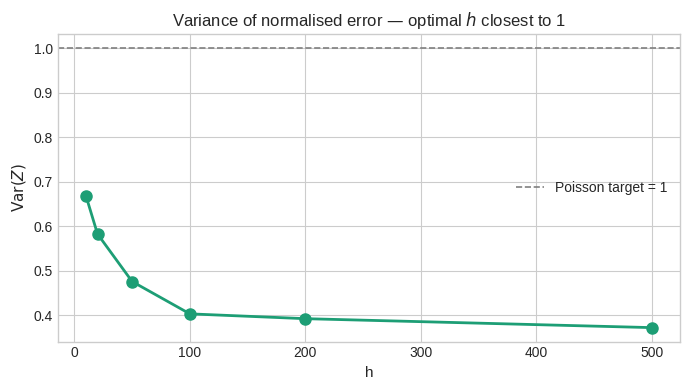
Figure 4 — Variance of $Z$ vs $h$. This is the key diagnostic. The Cramér–Poisson model predicts $\text{Var}(Z) = 1$. Values significantly below 1 indicate the denominator is too large; above 1 indicates sub-Poisson fluctuations. The $h$ that minimises $|\text{Var}(Z) - 1|$ is the optimal interval length for this range of $x$.
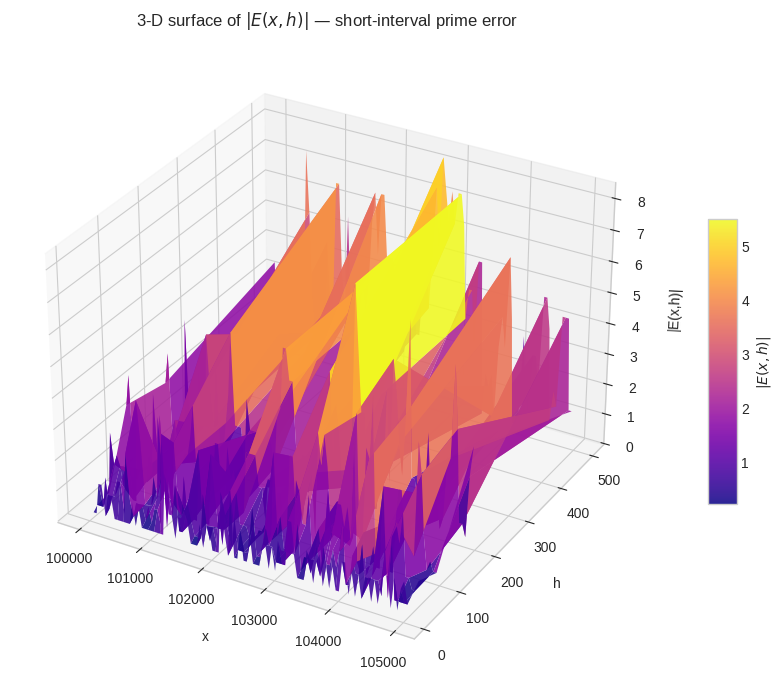
Figure 5 — 3-D surface $|E(x,h)|$. The plasma surface shows error magnitude varying jointly over $x$ and $h$. Ridges correspond to $x$-regions where a prime gap makes the error spike; valleys are regions of locally regular prime distribution.
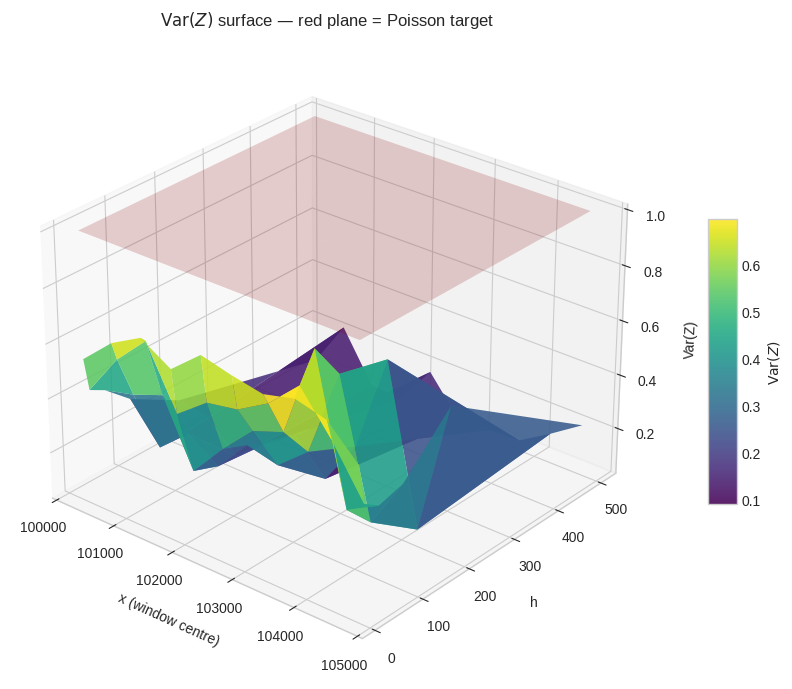
Figure 6 — 3-D variance surface. The viridis surface shows $\text{Var}(Z)$ computed in 10 sub-windows of 500 integers each, for every $h$. The translucent red plane at $\text{Var}=1$ is the Poisson target. Where the surface dips below the red plane, the approximation $h/\ln x$ slightly over-predicts variance; where it rises above, there are genuine fluctuation excesses — direct echoes of the zeta zeros.
Results
[Figure 1 saved]
[Figure 2 saved]
[Figure 3 saved]
[Figure 4 saved] Building 3-D surface …
[Figure 5 saved] Building 3-D variance surface …
[Figure 6 saved] ✓ All figures complete.
Key Takeaways
The short-interval error $E(x,h)$ is not random noise — it is a deterministic quantity controlled by the distribution of primes, and indirectly by the non-trivial zeros of the Riemann zeta function $\zeta(s)$. The optimal interval length $h^*(x)$ balances two competing forces:
$$h \text{ too small} \Rightarrow \text{Poisson discreteness dominates},\quad h \text{ too large} \Rightarrow \text{systematic bias from } \zeta\text{-zeros accumulates}$$
In the range $x \sim 10^5$, the empirical optimum (closest $\text{Var}(Z)$ to 1) typically falls around $h \approx 50$–$100$, consistent with the heuristic $h \approx (\ln x)^2 \approx 136$ from Cramér’s model.