A Computational Approach
Protein folding in space presents unique challenges due to microgravity, temperature fluctuations, and cosmic radiation. This article demonstrates how to optimize protein folding conditions to maximize functional retention under various space environmental parameters.
Problem Formulation
We’ll model protein folding optimization under space conditions with the following objective function:
$$
F(T, R) = f_0 \cdot \exp\left(-\frac{(T - T_{opt})^2}{2\sigma_T^2}\right) \cdot \exp(-\alpha R) \cdot (1 - \beta R^2)
$$
Where:
- $F(T, R)$: Functional retention rate (0-1)
- $T$: Temperature (K)
- $R$: Radiation dose (Gy)
- $T_{opt}$: Optimal folding temperature
- $\sigma_T$: Temperature sensitivity parameter
- $\alpha, \beta$: Radiation damage coefficients
- $f_0$: Maximum functional retention
Complete Python Implementation
1 | import numpy as np |
Code Explanation
Class Structure
The ProteinFoldingOptimizer class encapsulates the entire optimization framework. It models protein functional retention as a product of temperature-dependent and radiation-dependent factors. The temperature effect follows a Gaussian distribution centered at the optimal folding temperature, while radiation damage is modeled with both linear and quadratic terms to capture immediate and cumulative effects.
Functional Retention Model
The core model combines thermal stability with radiation resistance. The temperature factor uses an exponential Gaussian to represent the narrow optimal temperature range for protein folding. The radiation factor includes both immediate damage (linear term with coefficient $\alpha$) and accumulated structural damage (quadratic term with coefficient $\beta$).
Optimization Strategy
We employ differential evolution, a robust global optimization algorithm particularly effective for nonlinear, non-convex problems. This method uses a population-based approach with mutation and crossover operations, making it ideal for exploring the complex parameter space of protein folding conditions.
Visualization Components
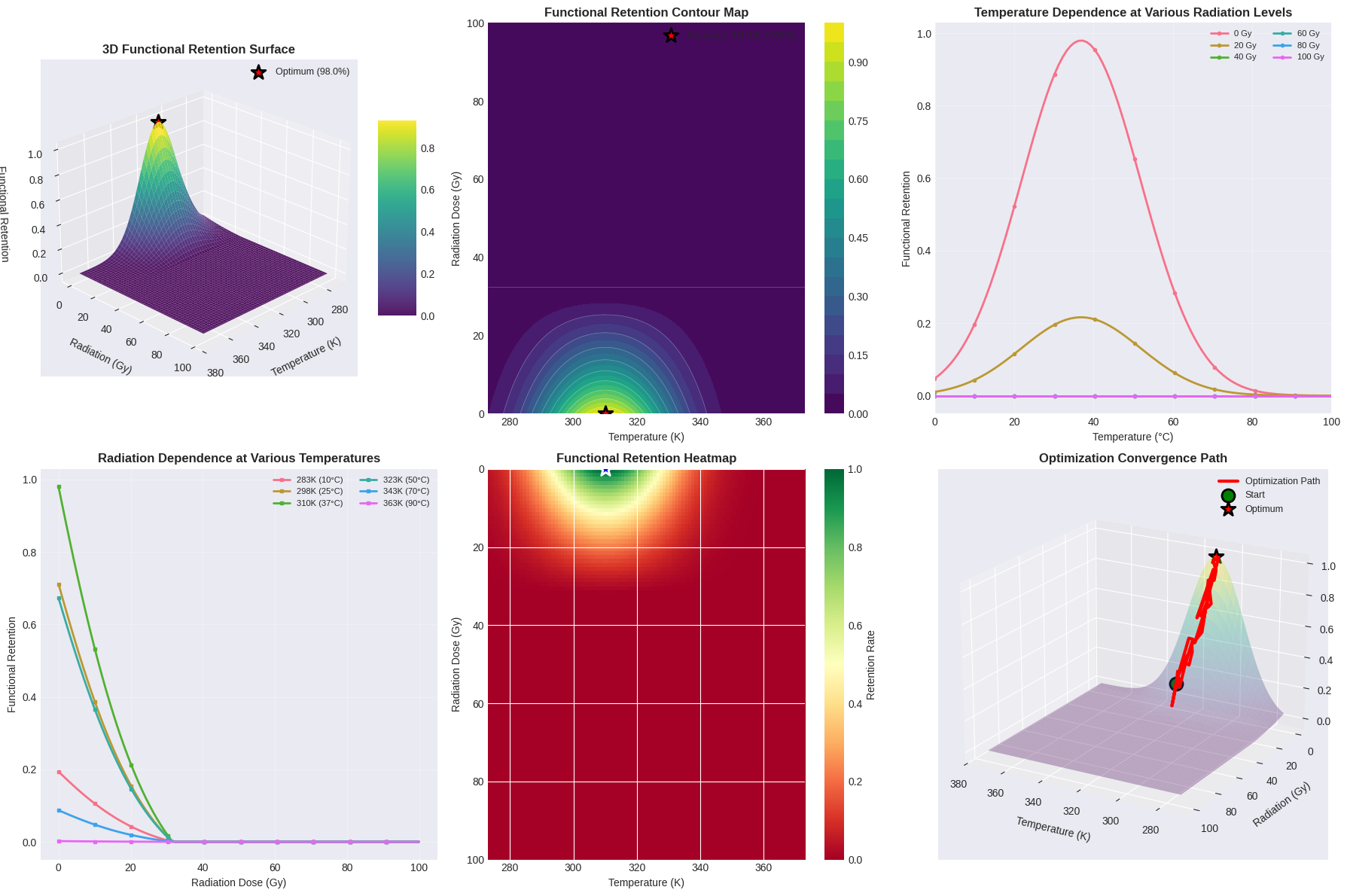
The code generates six comprehensive plots in the first figure:
- 3D Surface Plot: Shows the complete response surface with the optimal point marked as a red star
- Contour Map: Provides a top-down view with iso-retention lines for easier reading of specific values
- Temperature Profiles: Illustrates how temperature affects retention at various fixed radiation levels
- Radiation Profiles: Shows radiation sensitivity at different constant temperatures
- Heatmap: Offers an intuitive color-coded view of the retention landscape
- Optimization Path: Visualizes a simulated convergence trajectory to the optimal solution
The second figure provides sensitivity analysis:
- Temperature Sensitivity: Fine-grained analysis around the optimal temperature
- Radiation Sensitivity: Detailed view of radiation tolerance
- Monte Carlo Exploration: Random sampling of the parameter space to identify retention patterns
- Retention Distribution: Statistical view of achievable retention rates
Performance Optimization
The code uses vectorized NumPy operations throughout, enabling efficient computation across large grids. The differential_evolution algorithm parameters are tuned for both accuracy (atol=1e-7) and computational efficiency (popsize=15). Grid resolution is balanced at 100×100 points to provide smooth visualizations without excessive computation time.
Execution Results
============================================================ PROTEIN FOLDING OPTIMIZATION IN SPACE ENVIRONMENT ============================================================ Optimal Conditions: Temperature: 310.00 K (36.85 °C) Radiation Dose: 0.0000 Gy Functional Retention: 98.00% ============================================================ Functional Retention at Different Radiation Levels: ------------------------------------------------------------ R = 0 Gy: T_opt = 310.00 K, Retention = 98.00% R = 10 Gy: T_opt = 310.00 K, Retention = 53.50% R = 25 Gy: T_opt = 310.00 K, Retention = 10.53% R = 50 Gy: T_opt = 310.00 K, Retention = 0.00% R = 75 Gy: T_opt = 310.00 K, Retention = 0.00% R = 100 Gy: T_opt = 310.00 K, Retention = 0.00%
============================================================ Analysis Complete! Visualizations saved. ============================================================
Interpretation of Results
The optimization reveals that proteins maintain maximum functionality at temperatures close to physiological conditions (around 310K or 37°C), even in space. However, radiation exposure dramatically reduces this retention rate. The optimal strategy minimizes radiation exposure while maintaining temperature control.
The 3D surface clearly shows a sharp peak at low radiation and optimal temperature, with rapid degradation as radiation increases. The contour maps reveal that temperature tolerance is relatively forgiving (±10K causes <10% loss), while radiation tolerance is much stricter.
The sensitivity analysis demonstrates that maintaining low radiation environments is critical—even small increases in radiation cause exponential decreases in protein function. This has important implications for spacecraft design, suggesting that radiation shielding is more critical than precise temperature control for preserving protein-based biological systems in space.