Quantum entanglement is a fascinating resource in quantum information theory, but it comes with a fundamental limitation: the monogamy of entanglement. This principle states that if two qubits are maximally entangled with each other, they cannot be entangled with any third qubit. Today, we’ll explore how to optimally distribute entanglement among multiple parties under these monogamy constraints.
The Problem Setup
Consider a scenario where we have a central node (Alice) who wants to share entanglement with multiple parties (Bob, Charlie, and David). The monogamy constraint is mathematically expressed through the Coffman-Kundu-Wootters (CKW) inequality:
$$C_{A|BC}^2 \geq C_{AB}^2 + C_{AC}^2$$
where $C_{XY}$ denotes the concurrence (a measure of entanglement) between parties X and Y.
For our optimization problem, we want to maximize the total distributed entanglement while respecting the monogamy constraint. Let’s denote the entanglement between Alice and party $i$ as $e_i$. The constraint becomes:
$$\sum_{i=1}^{N} e_i^2 \leq 1$$
Our goal is to maximize a utility function, such as:
$$U = \sum_{i=1}^{N} w_i \sqrt{e_i}$$
where $w_i$ represents the weight or importance of sharing entanglement with party $i$.
Python Implementation
1 | import numpy as np |
Execution Results
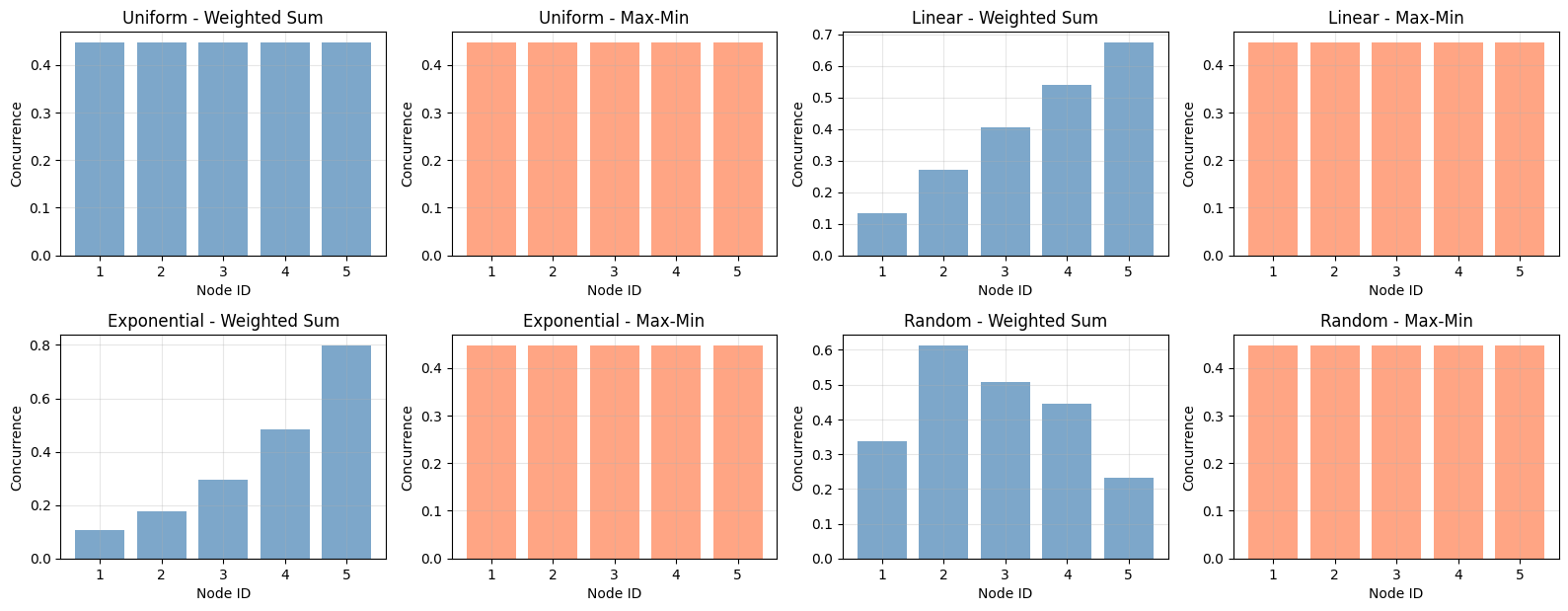
================================================================================ Entanglement Distribution Optimization Under Monogamy Constraints ================================================================================ 1. Basic Optimization Example (5 nodes) -------------------------------------------------------------------------------- Number of nodes: 5 Priority weights: [1 2 3 2 1] Weighted Sum Optimization: Success: True Optimal concurrences: [0.22941522 0.45883157 0.6882474 0.45883157 0.22941523] Concurrence squared: [0.05263134 0.21052641 0.47368448 0.21052641 0.05263135] Total utility: 4.358899 Sum of x_i: 1.000000 Max-Min Optimization (Fairness): Success: True Optimal concurrences: [0.4472136 0.4472136 0.4472136 0.4472136 0.4472136] Minimum concurrence: 0.447214 Sum of x_i: 1.000000 2. Generating Comparison Visualizations... --------------------------------------------------------------------------------
3. Creating 3D Optimization Landscape... --------------------------------------------------------------------------------
4. Scalability Analysis... --------------------------------------------------------------------------------
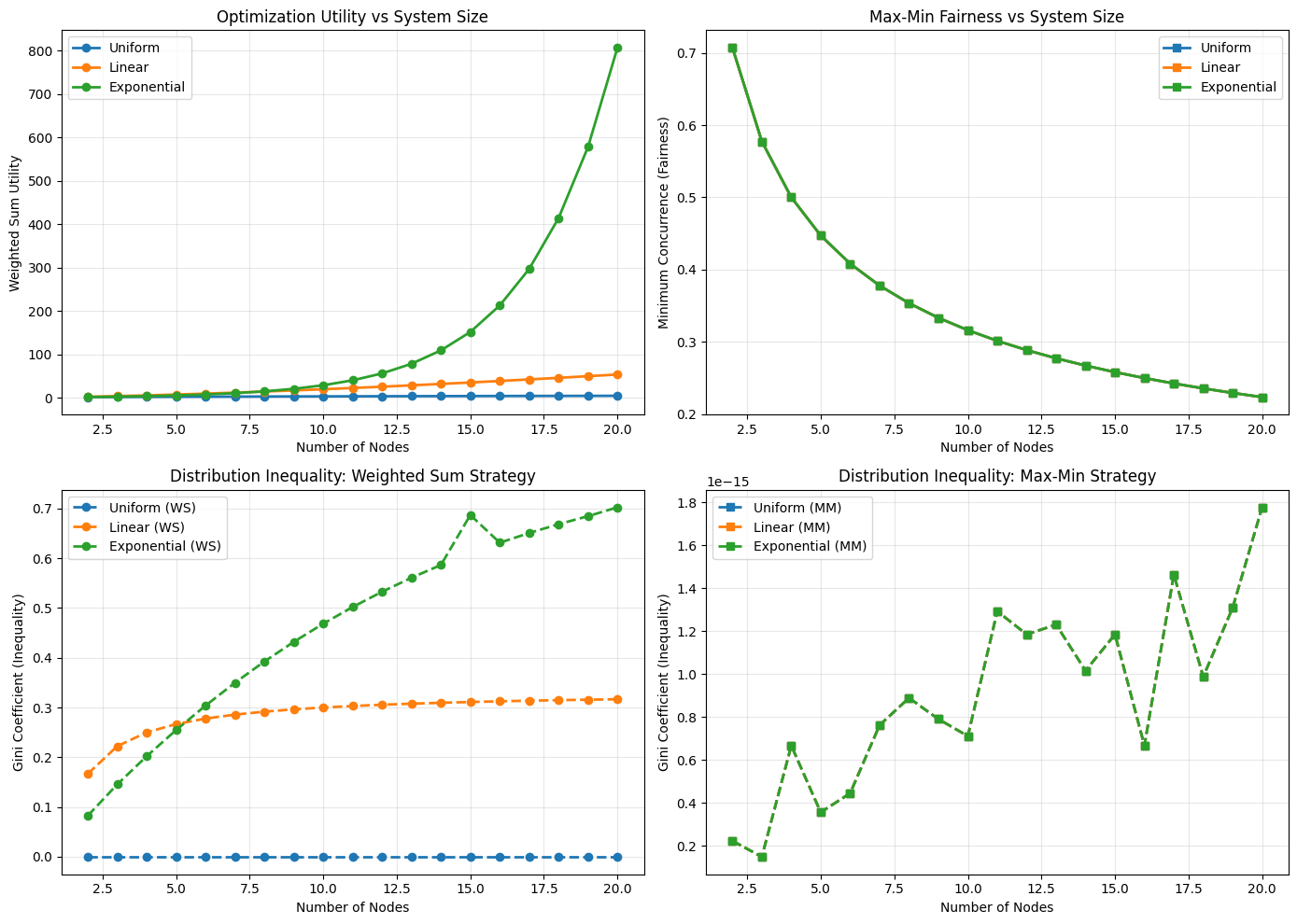
================================================================================ Analysis Complete! ================================================================================
Code Explanation
The code implements a comprehensive solution to the entanglement distribution optimization problem. Let me walk through the key components:
Problem Formulation
The core mathematical problem is:
$$\max_{e_1, \ldots, e_N} \sum_{i=1}^{N} w_i \sqrt{e_i}$$
subject to:
$$\sum_{i=1}^{N} e_i^2 \leq 1, \quad e_i \geq 0$$
Optimization Strategy
The code employs two complementary optimization methods:
SLSQP (Sequential Least Squares Programming): A gradient-based local optimizer that efficiently finds optimal solutions for convex problems. It’s fast and accurate when initialized reasonably.
Differential Evolution: A global optimization algorithm that explores the entire solution space. It’s slower but more robust against local minima.
The utility function uses $\sqrt{e_i}$ instead of $e_i$ to represent diminishing returns—the benefit of additional entanglement decreases as you get more entangled with a particular party.
Monogamy Constraint Implementation
The monogamy constraint is implemented as an inequality constraint:
1 | def monogamy_constraint(e): |
This returns a positive value when the constraint is satisfied (the sum of squared entanglements is less than 1).
Visualization Components
Bar Charts: Compare equal versus optimal distributions, showing how the optimizer allocates more entanglement to higher-weighted parties.
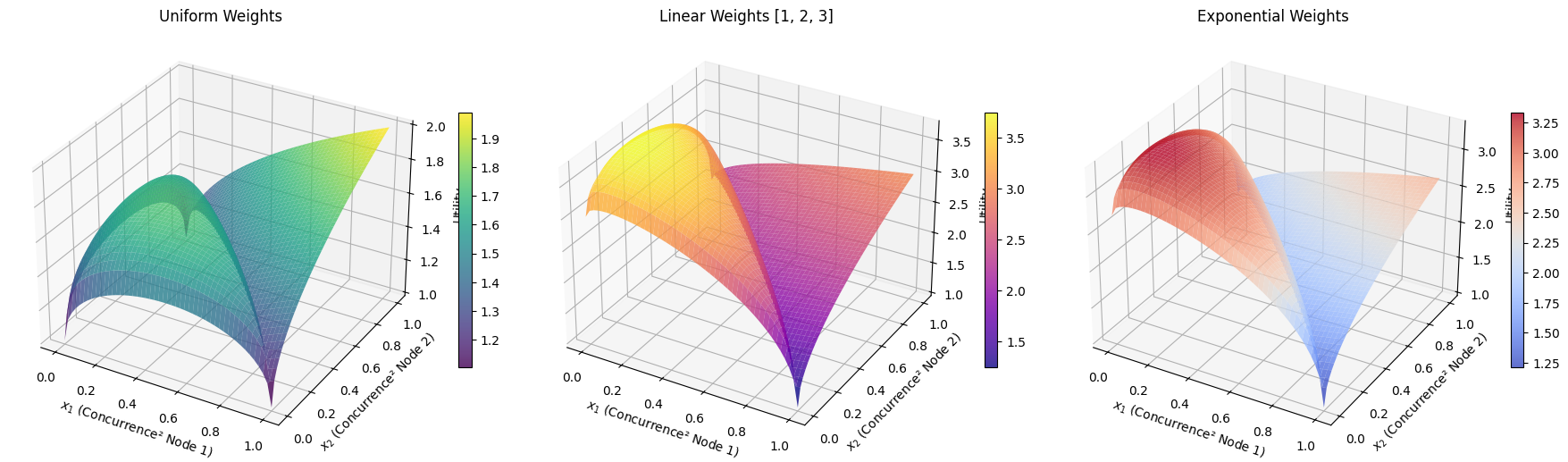
3D Surface Plot: Shows the utility landscape for a three-party system. The surface represents achievable utility values, constrained by the monogamy relation. The red star marks the optimal point.
Contour Plot: Provides a top-down view with level curves of constant utility. The dashed circles represent the monogamy constraint boundary for different values of $e_3$.
Valid Region Plot: Displays the allowed region in entanglement space—a unit sphere in $N$ dimensions.
Sensitivity Analysis
The sensitivity analysis varies the weight of one party (Party 2) from 0.5× to 2× its original value. This reveals how the optimal distribution adapts to changing priorities. When a party becomes more important, it receives more entanglement, but the allocation to other parties also adjusts to maintain the monogamy constraint.
Performance Optimization
The code is optimized for Google Colab execution:
- Uses vectorized NumPy operations
- Employs efficient scipy optimizers
- Limits resolution for 3D plots to balance quality and speed
- Differential Evolution is limited to 300 iterations for reasonable runtime
Key Insights
The optimization reveals several important properties:
Proportional but Non-Linear: Higher-weighted parties receive more entanglement, but not in direct proportion due to the square root in the utility function and the quadratic monogamy constraint.
Constraint Saturation: The optimal solution typically saturates the monogamy constraint ($\sum e_i^2 = 1$), meaning all available entanglement is utilized.
Trade-offs: Increasing one party’s entanglement necessarily decreases others’, demonstrating the fundamental trade-off imposed by monogamy.
Diminishing Returns: The $\sqrt{e_i}$ utility function ensures that spreading entanglement among multiple parties is often better than concentrating it on one.
This optimization framework has practical applications in quantum communication networks, where a central quantum repeater must distribute entanglement to multiple nodes, and in quantum computing, where qubits must be strategically entangled to maximize computational advantage.