1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
326
327
328
329
330
331
332
333
334
| import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import minimize_scalar
import seaborn as sns
plt.style.use('seaborn-v0_8')
sns.set_palette("husl")
def male_fitness(x, a=2.0, b=0.05, c=0):
"""
Male fitness function: F_m(x) = ax - bx^2 + c
Parameters:
- a: benefit coefficient from size (competition advantage)
- b: cost coefficient (metabolic cost increases quadratically)
- c: baseline fitness
"""
return a * x - b * x**2 + c
def female_fitness(x, d=1.5, e=0.03, f=0.001, g=0):
"""
Female fitness function: F_f(x) = dx - ex^2 - fx^3 + g
Parameters:
- d: benefit coefficient from size (resource acquisition)
- e: quadratic cost coefficient
- f: cubic cost coefficient (reproductive burden)
- g: baseline fitness
"""
return d * x - e * x**2 - f * x**3 + g
scenarios = {
"Scenario 1: Moderate dimorphism": {
"male_params": {"a": 2.0, "b": 0.05, "c": 0},
"female_params": {"d": 1.5, "e": 0.03, "f": 0.001, "g": 0}
},
"Scenario 2: Strong dimorphism": {
"male_params": {"a": 3.0, "b": 0.04, "c": 0},
"female_params": {"d": 1.2, "e": 0.06, "f": 0.002, "g": 0}
},
"Scenario 3: Weak dimorphism": {
"male_params": {"a": 1.8, "b": 0.06, "c": 0},
"female_params": {"d": 1.6, "e": 0.05, "f": 0.0005, "g": 0}
}
}
def find_optimal_sizes(male_params, female_params, x_range=(0, 50)):
"""
Find optimal body sizes for males and females using calculus-based optimization
"""
male_optimal = male_params["a"] / (2 * male_params["b"])
e, f, d = female_params["e"], female_params["f"], female_params["d"]
discriminant = 4 * e**2 + 12 * f * d
if discriminant >= 0 and f != 0:
female_optimal_1 = (-2 * e + np.sqrt(discriminant)) / (6 * f)
female_optimal_2 = (-2 * e - np.sqrt(discriminant)) / (6 * f)
candidates = [x for x in [female_optimal_1, female_optimal_2]
if x > 0 and x_range[0] <= x <= x_range[1]]
if candidates:
female_optimal = max(candidates)
else:
female_optimal = d / (2 * e)
else:
female_optimal = d / (2 * e)
return male_optimal, female_optimal
def calculate_dimorphism_index(male_size, female_size):
"""
Calculate sexual dimorphism index: (larger_sex_size - smaller_sex_size) / smaller_sex_size
"""
if male_size > female_size:
return (male_size - female_size) / female_size
else:
return (female_size - male_size) / male_size
results = {}
x_values = np.linspace(0, 50, 1000)
print("=== SEXUAL DIMORPHISM OPTIMIZATION ANALYSIS ===\n")
for scenario_name, params in scenarios.items():
male_params = params["male_params"]
female_params = params["female_params"]
male_fitness_values = [male_fitness(x, **male_params) for x in x_values]
female_fitness_values = [female_fitness(x, **female_params) for x in x_values]
male_optimal, female_optimal = find_optimal_sizes(male_params, female_params)
male_max_fitness = male_fitness(male_optimal, **male_params)
female_max_fitness = female_fitness(female_optimal, **female_params)
dimorphism_index = calculate_dimorphism_index(male_optimal, female_optimal)
results[scenario_name] = {
"x_values": x_values,
"male_fitness": male_fitness_values,
"female_fitness": female_fitness_values,
"male_optimal": male_optimal,
"female_optimal": female_optimal,
"male_max_fitness": male_max_fitness,
"female_max_fitness": female_max_fitness,
"dimorphism_index": dimorphism_index
}
print(f"--- {scenario_name} ---")
print(f"Optimal male body size: {male_optimal:.2f} units")
print(f"Optimal female body size: {female_optimal:.2f} units")
print(f"Male maximum fitness: {male_max_fitness:.3f}")
print(f"Female maximum fitness: {female_max_fitness:.3f}")
print(f"Sexual dimorphism index: {dimorphism_index:.3f}")
if male_optimal > female_optimal:
print(f"Males are {((male_optimal/female_optimal - 1) * 100):.1f}% larger than females")
else:
print(f"Females are {((female_optimal/male_optimal - 1) * 100):.1f}% larger than males")
print()
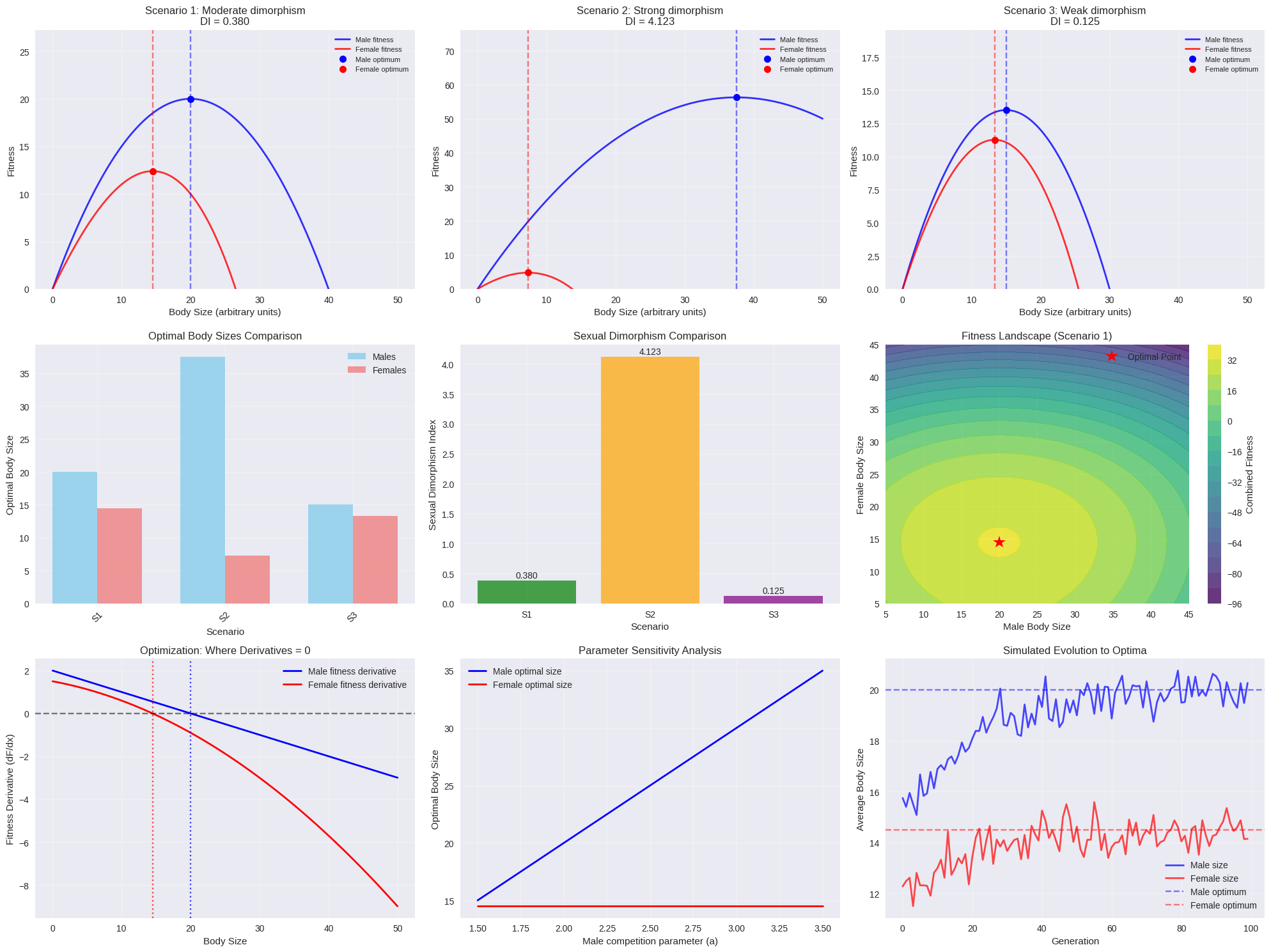
fig = plt.figure(figsize=(20, 15))
for i, (scenario_name, data) in enumerate(results.items()):
plt.subplot(3, 3, i+1)
plt.plot(data["x_values"], data["male_fitness"], 'b-', linewidth=2,
label=f'Male fitness', alpha=0.8)
plt.plot(data["x_values"], data["female_fitness"], 'r-', linewidth=2,
label=f'Female fitness', alpha=0.8)
plt.plot(data["male_optimal"], data["male_max_fitness"], 'bo',
markersize=8, label=f'Male optimum')
plt.plot(data["female_optimal"], data["female_max_fitness"], 'ro',
markersize=8, label=f'Female optimum')
plt.axvline(data["male_optimal"], color='blue', linestyle='--', alpha=0.5)
plt.axvline(data["female_optimal"], color='red', linestyle='--', alpha=0.5)
plt.xlabel('Body Size (arbitrary units)')
plt.ylabel('Fitness')
plt.title(f'{scenario_name}\nDI = {data["dimorphism_index"]:.3f}')
plt.legend(fontsize=8)
plt.grid(True, alpha=0.3)
plt.ylim(bottom=0)
plt.subplot(3, 3, 4)
scenario_names = list(results.keys())
male_sizes = [results[s]["male_optimal"] for s in scenario_names]
female_sizes = [results[s]["female_optimal"] for s in scenario_names]
x_pos = np.arange(len(scenario_names))
width = 0.35
plt.bar(x_pos - width/2, male_sizes, width, label='Males', color='skyblue', alpha=0.8)
plt.bar(x_pos + width/2, female_sizes, width, label='Females', color='lightcoral', alpha=0.8)
plt.xlabel('Scenario')
plt.ylabel('Optimal Body Size')
plt.title('Optimal Body Sizes Comparison')
plt.xticks(x_pos, [f'S{i+1}' for i in range(len(scenario_names))], rotation=45)
plt.legend()
plt.grid(True, alpha=0.3)
plt.subplot(3, 3, 5)
dimorphism_indices = [results[s]["dimorphism_index"] for s in scenario_names]
colors = ['green', 'orange', 'purple']
bars = plt.bar(range(len(scenario_names)), dimorphism_indices,
color=colors, alpha=0.7)
plt.xlabel('Scenario')
plt.ylabel('Sexual Dimorphism Index')
plt.title('Sexual Dimorphism Comparison')
plt.xticks(range(len(scenario_names)), [f'S{i+1}' for i in range(len(scenario_names))])
plt.grid(True, alpha=0.3)
for bar, value in zip(bars, dimorphism_indices):
plt.text(bar.get_x() + bar.get_width()/2, bar.get_height() + 0.01,
f'{value:.3f}', ha='center', va='bottom')
plt.subplot(3, 3, 6)
scenario_data = results[list(results.keys())[0]]
male_sizes_2d = np.linspace(5, 45, 50)
female_sizes_2d = np.linspace(5, 45, 50)
X, Y = np.meshgrid(male_sizes_2d, female_sizes_2d)
Z = np.zeros_like(X)
for i in range(X.shape[0]):
for j in range(X.shape[1]):
male_fit = male_fitness(X[i,j], **scenarios[list(scenarios.keys())[0]]["male_params"])
female_fit = female_fitness(Y[i,j], **scenarios[list(scenarios.keys())[0]]["female_params"])
Z[i,j] = male_fit + female_fit
contour = plt.contourf(X, Y, Z, levels=20, cmap='viridis', alpha=0.8)
plt.colorbar(contour, label='Combined Fitness')
plt.xlabel('Male Body Size')
plt.ylabel('Female Body Size')
plt.title('Fitness Landscape (Scenario 1)')
opt_data = results[list(results.keys())[0]]
plt.plot(opt_data["male_optimal"], opt_data["female_optimal"], 'r*',
markersize=15, label='Optimal Point')
plt.legend()
plt.subplot(3, 3, 7)
scenario_data = results[list(results.keys())[0]]
x_vals = scenario_data["x_values"]
male_derivative = np.gradient(scenario_data["male_fitness"], x_vals)
female_derivative = np.gradient(scenario_data["female_fitness"], x_vals)
plt.plot(x_vals, male_derivative, 'b-', linewidth=2, label="Male fitness derivative")
plt.plot(x_vals, female_derivative, 'r-', linewidth=2, label="Female fitness derivative")
plt.axhline(y=0, color='k', linestyle='--', alpha=0.5)
plt.axvline(scenario_data["male_optimal"], color='blue', linestyle=':', alpha=0.7)
plt.axvline(scenario_data["female_optimal"], color='red', linestyle=':', alpha=0.7)
plt.xlabel('Body Size')
plt.ylabel('Fitness Derivative (dF/dx)')
plt.title('Optimization: Where Derivatives = 0')
plt.legend()
plt.grid(True, alpha=0.3)
plt.subplot(3, 3, 8)
a_values = np.linspace(1.5, 3.5, 20)
male_optima = []
female_optima = []
base_male_params = {"a": 2.0, "b": 0.05, "c": 0}
base_female_params = {"d": 1.5, "e": 0.03, "f": 0.001, "g": 0}
for a_val in a_values:
male_params = base_male_params.copy()
male_params["a"] = a_val
male_opt, female_opt = find_optimal_sizes(male_params, base_female_params)
male_optima.append(male_opt)
female_optima.append(female_opt)
plt.plot(a_values, male_optima, 'b-', linewidth=2, label='Male optimal size')
plt.plot(a_values, female_optima, 'r-', linewidth=2, label='Female optimal size')
plt.xlabel('Male competition parameter (a)')
plt.ylabel('Optimal Body Size')
plt.title('Parameter Sensitivity Analysis')
plt.legend()
plt.grid(True, alpha=0.3)
plt.subplot(3, 3, 9)
generations = np.arange(0, 100)
male_size_evolution = []
female_size_evolution = []
initial_male_size = 15
initial_female_size = 12
target_male = scenario_data["male_optimal"]
target_female = scenario_data["female_optimal"]
for gen in generations:
progress = 1 - np.exp(-gen/20)
current_male = initial_male_size + (target_male - initial_male_size) * progress
current_female = initial_female_size + (target_female - initial_female_size) * progress
current_male += np.random.normal(0, 0.5)
current_female += np.random.normal(0, 0.5)
male_size_evolution.append(current_male)
female_size_evolution.append(current_female)
plt.plot(generations, male_size_evolution, 'b-', linewidth=2, alpha=0.7, label='Male size')
plt.plot(generations, female_size_evolution, 'r-', linewidth=2, alpha=0.7, label='Female size')
plt.axhline(target_male, color='blue', linestyle='--', alpha=0.5, label='Male optimum')
plt.axhline(target_female, color='red', linestyle='--', alpha=0.5, label='Female optimum')
plt.xlabel('Generation')
plt.ylabel('Average Body Size')
plt.title('Simulated Evolution to Optima')
plt.legend()
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print("\n=== SUMMARY STATISTICS ===")
all_male_sizes = [results[s]["male_optimal"] for s in results.keys()]
all_female_sizes = [results[s]["female_optimal"] for s in results.keys()]
all_dimorphism = [results[s]["dimorphism_index"] for s in results.keys()]
print(f"Average optimal male size: {np.mean(all_male_sizes):.2f} ± {np.std(all_male_sizes):.2f}")
print(f"Average optimal female size: {np.mean(all_female_sizes):.2f} ± {np.std(all_female_sizes):.2f}")
print(f"Average dimorphism index: {np.mean(all_dimorphism):.3f} ± {np.std(all_dimorphism):.3f}")
print(f"Range of dimorphism: {min(all_dimorphism):.3f} - {max(all_dimorphism):.3f}")
print("\n=== MATHEMATICAL FORMULAS USED ===")
print("Male fitness function: F_m(x) = ax - bx² + c")
print("Female fitness function: F_f(x) = dx - ex² - fx³ + g")
print("Male optimum: x* = a/(2b)")
print("Female optimum: x* = solution to 3fx² + 2ex - d = 0")
print("Sexual Dimorphism Index: (|size_male - size_female|) / min(size_male, size_female)")
|